The Boston Children’s Hospital’s science and clinical innovation blog Vector (February 2019 by Nancy Fliesler), features the new paper titled “NMDA 2A receptors in parvalbumin cells mediate sex-specific rapid ketamine response on cortical activity”.
Read the Vector blog entitled “How the antidepressant ketamine rapidly awakens the brain and why its effects vary more in women”
Geographical maps have long changed the way we live, from the ancient hand-drawn ones that relied on explorer voyages to those on today’s satellite-driven, real time apps like Google Maps. The same transformative power holds true for maps in neuroscience. From the meticulous illustrations of neurons in different brain regions made by neuroanatomist Ramon y Cajal in the late 1800s to the imaging of neural circuits through more modern computerized technologies, the maps that we have of our nervous systems are constantly shaping the questions we think to ask and are able to answer about how our brains and bodies work—and what to do when diseases or injuries arise.
A recent advance in the map-making world of brain scientists comes from the lab of Xiaowei Zhuang, a faculty member in the Conte Center at Harvard who is the David B. Arnold Professor of Science and a Howard Hughes Medical Institute Investigator. In a study published in Cell, led by postdoctoral fellows Yaron Sigal and Colenso Speer, her team showcases a powerful new approach for visualizing and analyzing the input fields of neurons, using neurons in the eye called retinal ganglion cells as a test case. While mapping the sea of inputs that a single neuron receives and integrates is something that neuroscientists have long recognized as critical for understanding the functions of different kinds of neurons and their role in circuits—and have been able to accomplish in varying capacities—until now there’s usually been a trade-off between obtaining very high resolution images and having precise molecular information. The Zhuang team’s new strategy will allow researchers automated analysis of neuron input fields in a way that minimizes this compromise and overlays complex molecular information upon three dimensional reconstructions of neurons situated in relatively thick sections of brain tissue.
When it comes to science, perhaps the wisest amongst us are those who know how much we don’t know. The famous 19th century neuroanatomist Santiago Ramón y Cajal, who many would argue did more to characterize the different cell types that make up the nervous system than any other scientist of his time, once described the adult brain as a ‘jungle’ that is ‘impenetrable and indefinable.’
Over a hundred years later, despite an explosion of new technologies in microscopy and genetic and immunological methods of brain cell labeling, neuroscience research remains chock full of uncharted territories. In a recent publication in Cell, a team led by Jeff Lichtman, the Jeremy R. Knowles Professor of Molecular and Cellular Biology at Harvard—fittingly also named the Santiago Ramón y Cajal Professor of Arts and Sciences—presents the latest chapter in this story, by providing unparalleled views into the intricacies of synaptic connectivity in the cerebral cortex and revealing novel insights about this connectivity—while simultaneously highlighting just how much we have left to discover.
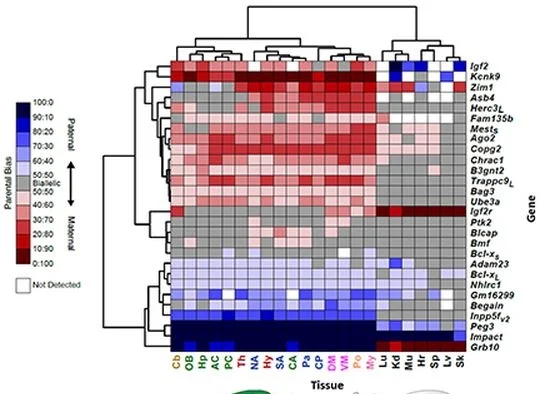
Original Research Article in eLife:
Quantitative and functional interrogation of parent-of-origin allelic expression biases in the brain.
By Perez, Rubinstein et al. July 3, 2015.
Harvard Gazette: Expanding the Brain. By Peter Reuell, July 29, 2015.
A new review explains how different methods of tissue clearing work
Imaging biological tissues is always a challenge because they are three dimensional, and issues of light scatter make it hard to clearly view deeper structures without cutting sections. Neuroscientists have long been interested in methods for “clearing” - or making more transparent -brain tissue so it is easier to image intact.
“I think as a neuroscientist there’s no part of the body where clearing is potentially more influential in moving forward,” says Jeff Lichtman, a professor of molecular and cellular biology and Conte lab head who teaches microscopy at Harvard and the Marine Biological Laboratory in Woods Hole.
Aired May 25th 2015. Features Catherine Dulac of our center in addition to Eric Kandel, Susanne Shultz, Charles Nelson, Margaret Spinelli and David Levine.
Aired June 16th 2015. Features Catherine Dulac of our center in addition to Eric Kandel, Ben Barres, Norman Spack, Melissa Hines and Janet Hyde.
Harvard Medical School News: FISHing For Insight >>
Story by Stephanie Dutchen, May 21st. New method for visualizing chromosomes in astonishing detail, developed by Ting Wu's lab in collaboration with Xiawei Zhuang and Peng Yin's groups, published May 12 in Nature Communications.
BBC World Service:The Forum – Plasticity >>
Aired May 19th. Features Takao Hensch of our center talking about brain plasticity alongside an artist who works with plastics from garbage and a nanochemist who considers plasticity at the level of atoms and electrons. Host Bridget Kendall.
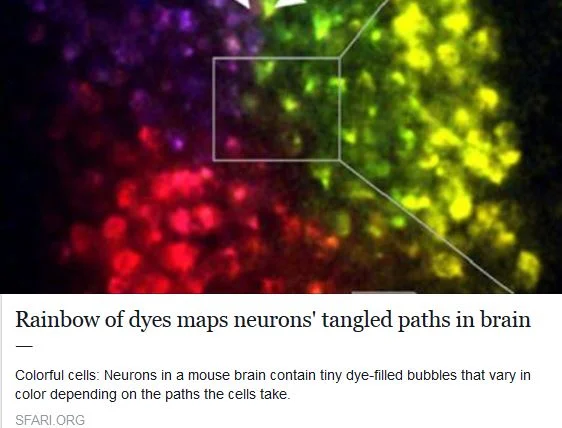
SFARI Autism News: Rainbow Of Dyes Maps Neurons’ Tangled Paths In Brain >>
Story by Kate Yandell, May 13th. Discusses neuronal positioning system (NPS) recently developed by Jeff Lichtman's lab, published April 21 in Nature Methods.